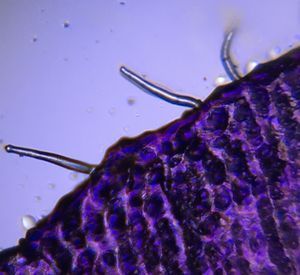
A simple microfluidic device and time lapse imaging of embryonic stem cells
 Mar 04, 2016 • 9:07 AM UTC
Mar 04, 2016 • 9:07 AM UTC Unknown Location
Unknown Location 140x Magnification
140x Magnification Microorganisms
Microorganisms
Josh Guild
Learn about the author...
5posts
2comments
1locations

View in Media Gallery
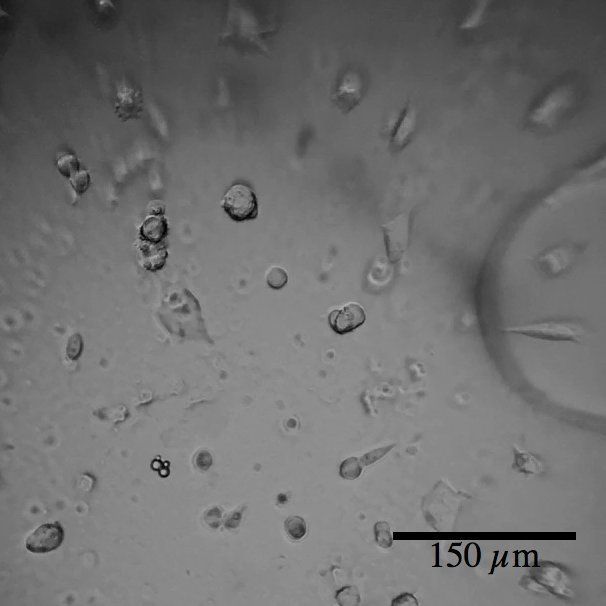
One thing that is overlooked in our general understanding of cell biology is how dynamic everything is. During school, you are taught that cells are a tiny round blobs full of even tinier factories performing various pre-defined functions. These cells are each of a certain type and shape and combine to form greater structures that do higher-level functions, such as transmit information over long distances or contract to generate force. When I see cells I see something that is much different. They do not form single defined shapes; they are dynamic and ever changing. They grow and shrink, change shape and move around. They reach out with small arm-like protrusions. They react and respond to their environment. They even can change their identity entirely.
We as humans are a fascinating example of all of these characteristics. The body begins as a single cell that divides to create many more of itself. As these cells divide they communicate with each other, change into other cell types, and move and organize themselves. Eventually they become a person; made up of trillions of cells categorized into over 200 types. Embryonic stem cells represent the earliest cells from which the body forms and are thus considered pluripotent, meaning that they have the potential to become any cell type in the body. Mouse embryonic stem cells (mESCs) are similar to their human counterparts in many regards and are commonly used as model to study these characteristics.
mESCs can be grown in the lab indefinitely. They divide about once every 16 hours and when they get too crowded, we take them and split them onto new dishes. Then, when needed, we can change their environment in ways that lead them to become other cell types, such as heart muscle or the neurons that make up the brain and spinal cord.
Using the Foldscope, my iPhone, a small microfluidic chamber for them to grow in, and an app that allowed me to snap one image every minute or two, I was able to watch them in action.
Fabrication of the microfluidic device
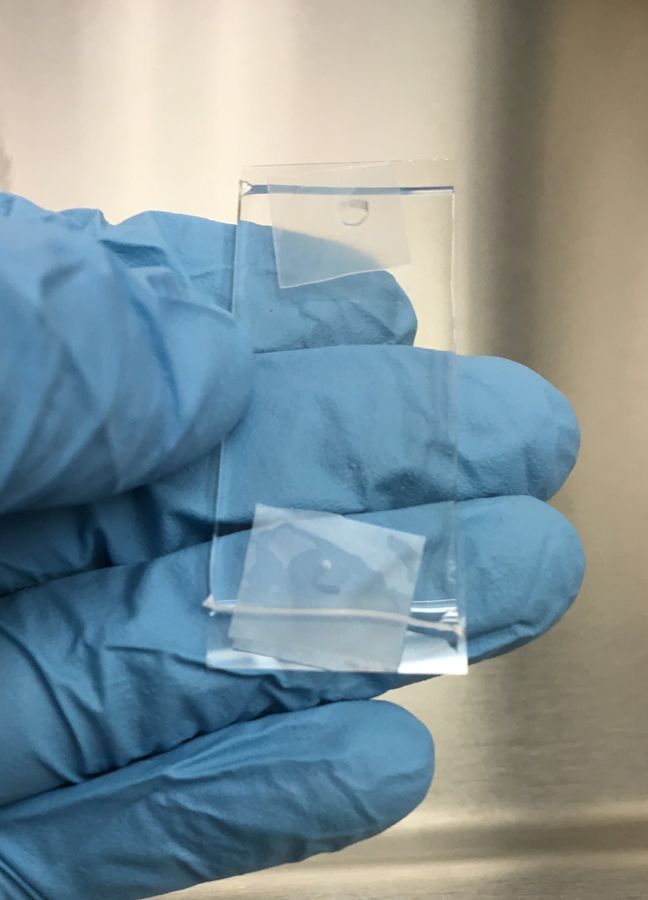
To image with the Foldscope, I used a microfluidic device that creates a protected enclosure to put cells in and that can be mounted on the Foldscope. The sealed chamber affords space in all directions for cells to proliferate and to be bathed in their required nutrient rich media.
The chamber is created out of a PDMS (polydimethylsiloxane) polymer (10 parts pre-polymer mixed thoroughly with 1 part curing reagent) that is degassed and poured onto a micro-fabricated negative relief of the cell culture chamber (20 mm length x 16 mm width x 60 µm height with small inlet and outlet channels to allow entry of cells and media exchange). The PDMS is then solidified by baking at 80C for 2 hours, pealed off of the mold, and attached to a glass slide through a process called oxygen plasma cleaning. (The roof of this design happened to also have 200 µm diameter pillars that served as nice way to generate a rough scale bar).
We as humans are a fascinating example of all of these characteristics. The body begins as a single cell that divides to create many more of itself. As these cells divide they communicate with each other, change into other cell types, and move and organize themselves. Eventually they become a person; made up of trillions of cells categorized into over 200 types. Embryonic stem cells represent the earliest cells from which the body forms and are thus considered pluripotent, meaning that they have the potential to become any cell type in the body. Mouse embryonic stem cells (mESCs) are similar to their human counterparts in many regards and are commonly used as model to study these characteristics.
mESCs can be grown in the lab indefinitely. They divide about once every 16 hours and when they get too crowded, we take them and split them onto new dishes. Then, when needed, we can change their environment in ways that lead them to become other cell types, such as heart muscle or the neurons that make up the brain and spinal cord.
Using the Foldscope, my iPhone, a small microfluidic chamber for them to grow in, and an app that allowed me to snap one image every minute or two, I was able to watch them in action.
Fabrication of the microfluidic device
To image with the Foldscope, I used a microfluidic device that creates a protected enclosure to put cells in and that can be mounted on the Foldscope. The sealed chamber affords space in all directions for cells to proliferate and to be bathed in their required nutrient rich media.
The chamber is created out of a PDMS (polydimethylsiloxane) polymer (10 parts pre-polymer mixed thoroughly with 1 part curing reagent) that is degassed and poured onto a micro-fabricated negative relief of the cell culture chamber (20 mm length x 16 mm width x 60 µm height with small inlet and outlet channels to allow entry of cells and media exchange). The PDMS is then solidified by baking at 80C for 2 hours, pealed off of the mold, and attached to a glass slide through a process called oxygen plasma cleaning. (The roof of this design happened to also have 200 µm diameter pillars that served as nice way to generate a rough scale bar).

View in Media Gallery
Seeding cells in the culture chamber
Cells attach to a class of proteins that together make up what is called the extracellular matrix (ECM). The ECM sticks to and coats the surrounding area, and provides a structural support for the cells to grip onto. mESCs are best cultured when attach to gelatin coated surfaces. To coat the chambers I incubated overnight at 4C with a 0.2% gelatin solution.
The next day I rinsed out the excess gelatin, and flowed in fresh culture media (composed of a complex mixture of various nutrients and growth factors). I then added 40 µL of cells to the inlet, suspended in the same media at a concentration of 4 million cells/mL, and gently tilted the device to allow gravity to guide the cells to their desired location. Next, I placed parafilm over the inlet and outlet channel openings to prevent evaporation and incubated at 37C for two hours to allow the cells to adhere to the gelatin-coated surface.
Cells attach to a class of proteins that together make up what is called the extracellular matrix (ECM). The ECM sticks to and coats the surrounding area, and provides a structural support for the cells to grip onto. mESCs are best cultured when attach to gelatin coated surfaces. To coat the chambers I incubated overnight at 4C with a 0.2% gelatin solution.
The next day I rinsed out the excess gelatin, and flowed in fresh culture media (composed of a complex mixture of various nutrients and growth factors). I then added 40 µL of cells to the inlet, suspended in the same media at a concentration of 4 million cells/mL, and gently tilted the device to allow gravity to guide the cells to their desired location. Next, I placed parafilm over the inlet and outlet channel openings to prevent evaporation and incubated at 37C for two hours to allow the cells to adhere to the gelatin-coated surface.


After the cells attached I sealed the inlet and outlet with scotch tape, inverted the device, inserted it into the Foldscope, and was ready to image. I sealed the Foldscope with my phone attached in a sterile container and placed it in an incubator set at 37C.

View in Media Gallery
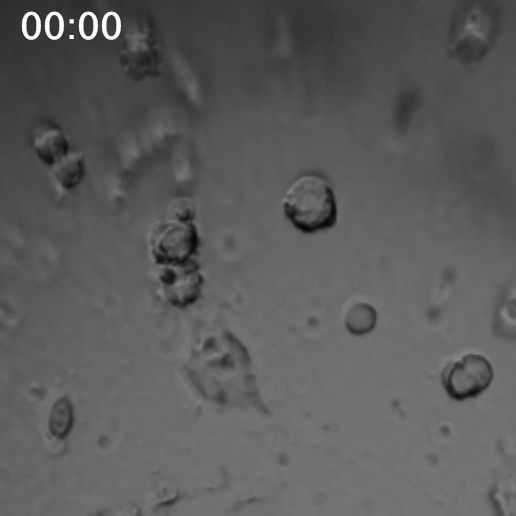
The first video was shot over an hour and 40 minutes with a photo taken every two minutes.

View in Media Gallery
The second video below was shot over 40 minutes with a photo taken every minute, and with slightly higher magnification. What is fascinating in this next video to me is watching what I thought was a single cell, just below and to the left of the center, jump apart into two cells and then come back together.

View in Media Gallery
DOI: 10.15200/winn.145765.50628 provided by The Winnower , a DIY scholarly publishing platform
Sign in to commentNobody has commented yet... Share your thoughts with the author and start the discussion!

 0 Applause
0 Applause 0 Comments
0 Comments